MassFlow MS Data Structures
This document describes the core data structures used in MassFlow for mass spectrometry (MS) workflows, with focus on loading and organizing spectra from ImzML. It covers the following modules and their relationships:
module/ms_module.pymodule/ms_data_manager.pymodule/ms_data_manager_imzml.py
The intent is to present implementation-focused details about classes, attributes, indexing patterns, and the lazy-loading data flow.
Overview
MassFlow separates domain models (data structures) from data managers (I/O and orchestration):
- Domain model
SpectrumBaseModule: Represents a single spectrum with spatial coordinates.SpectrumImzML: Specialized spectrum class for lazy-loading data from ImzML.MS: Collection of spectra with efficient coordinate-based indexing.
- Data manager
MSDataManager: Abstract base class defining common options and interface.MSDataManagerImzML: Concrete manager that reads.imzMLfiles and populatesMS.
Metadata is handled separately (see module/meta_data.py) and attached to managers/models as needed.
Class Diagram (Inheritance)
The following diagram shows the inheritance relationships among the core classes described above. SpectrumImzML extends SpectrumBaseModule, and MSDataManagerImzML extends MSDataManager. MS is a standalone container class with no inheritance.
The MSDataManager and its concrete implementations (like MSDataManagerImzML) are responsible for populating the MS container with spectrum data. The manager holds a reference to an MS instance and loads spectra into it through the load_full_data_from_file() method. The MS container holds multiple SpectrumBaseModule instances (or its subclasses like SpectrumImzML), organizing them by spatial coordinates for efficient access.
Core Types
SpectrumBaseModule Class
class module.ms_module.SpectrumBaseModule(mz_list, intensity, coordinates, sorted_by_mz_fun=False)Represents a single mass spectrum bound to spatial coordinates. Key characteristics:
Parameters
mz_list(Optional[np.ndarray]) — Array of m/z values. Can beNonefor lazy loading.intensity(Optional[np.ndarray]) — Array of intensity values. Can beNonefor lazy loading.coordinates(List[int]) — List of three integers[x, y, z]representing spatial coordinates.sorted_by_mz_fun(bool, optional) — Whether the data is already sorted by m/z. Defaults toFalse.
Properties
mz_list: np.ndarray— getter/setter; may beNoneuntil loaded or set.intensity: np.ndarray— getter/setter; may beNoneuntil loaded or set.
Utilities
sort_by_mz()— optional in-place sorting of spectrum by m/z.plot(...)— convenience plotting withlineorstemmodes.
Invariants and notes
coordinatesmust be a list of exactly 3 integers; indexing assumes[x, y, z]order.mz_listandintensitymust be same length when both are present.- Lazy loading is supported via
Noneinitialization, deferring data acquisition.
Example
>>> from tools.plot import plot_spectrum
>>> spectrum = ms[0] or ms.get_spectrum(0, 0, 0)
>>> mz_list = spectrum.mz_list
>>> print(mz_list)
output:
[16441.998 938.1308 2318.6423 ... 1174.1575 1333.138 1488.291 ]
>>> # Plot the spectrum using the standalone plotting utility
>>> plot_spectrum(base=spectrum)- Example2
if __name__ == "__main__":
from module.ms_data_manager_imzml import MSDataManagerImzML
from module.ms_module import MS
from tools.plot import plot_spectrum
FILE_PATH = "data/example.imzML"
ms = MS()
# Create MS collection and manager
# Auto-created from parser
with MSDataManagerImzML(ms=ms,target_locs=[(1, 1), (50, 50)],filepath=FILE_PATH) as manager:
# Load data with lazy-loading placeholders
manager.load_full_data_from_file()
spectrum = ms[0]
plot_spectrum(spectrum)
output: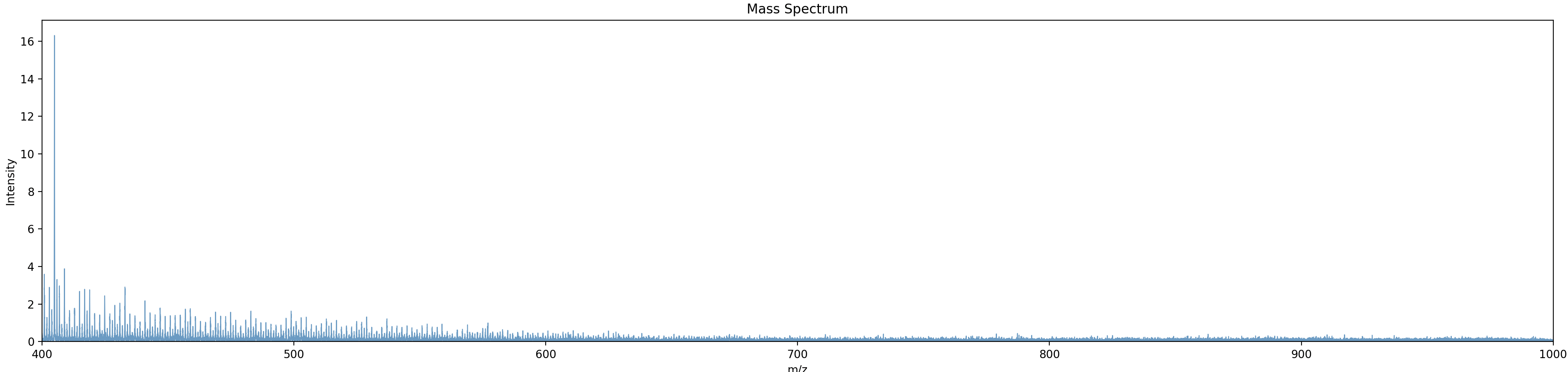
SpectrumImzML Class
class module.ms_module.SpectrumImzML(parser, index, coordinates)Specialized mass spectrum class for ImzML format with lazy loading capabilities. This class extends SpectrumBaseModule to provide efficient handling of ImzML (Imaging Mass Spectrometry Markup Language) format data by implementing lazy loading to minimize memory usage.
Note: use MSDataManagerImzML to automatically create and manage SpectrumImzML.
- Parameters
parser(ImzMLParser) — ImzML parser instance for reading spectrum data from the file.index(int) — Index of the spectrum within the ImzML file.coordinates(List[int]) — List of three integers[x, y, z]representing spatial coordinates of the spectrum location.
- Inherited Attributes
coordinates(List[int]) — 3D coordinates[x, y, z]of the spectrum.x,y,z(int) — Individual coordinate components.sorted_by_mz_fun(bool) — Flag indicating if data is sorted by m/z values.
- Properties
mz_list: np.ndarray— Lazily loaded array of m/z values; triggers data loading on first access.intensity: np.ndarray— Lazily loaded array of intensity values; ensures data is loaded when accessed.
- Lazy loading behavior
- On first
mz_listaccess, callsparser.getspectrum(index)to load both m/z and intensity data together. intensityaccess triggersmz_listloading if intensity is not yet available, ensuring data consistency.- Data loading is deferred until first property access to minimize memory footprint.
- On first
- Invariants and notes
- The actual m/z and intensity data are not loaded during initialization.
- Both
mz_listandintensityare loaded together for efficiency when either is first accessed. - Inherits all visualization and manipulation methods from
SpectrumBaseModule.
- Example
>>> # Create SpectrumImzML instance (data not yet loaded)
>>> spectrum = SpectrumImzML(parser, index=0, coordinates=[0, 0, 0])
>>> # First access triggers lazy loading
>>> mz_values = spectrum.mz_list
>>> print(mz_values[:5])
output:
[100.05 150.12 200.34 250.67 300.89]MS Class
class module.ms_module.MS()Collection class for managing multiple mass spectra with coordinate-based indexing. This class serves as a container and manager for multiple SpectrumBaseModule instances, providing efficient storage, retrieval, and manipulation of mass spectrometry data organized by 3D spatial coordinates.
- Parameters
- No parameters required for initialization.
- Attributes
meta(Optional) — Metadata object associated with the spectrum collection.
- Methods
add_spectrum(spectrum)— Add a mass spectrum to the collection with automatic coordinate indexing.get_spectrum(x, y, z=0)— Retrieve a mass spectrum by its 3D coordinates.plot_ms(x, y, z, save_path, figsize, dpi, color, plot_mode)— Plot a mass spectrum at specified coordinates.
- Indexing patterns
ms[index]— Sequential access by integer index into_queue.ms[x, y]— 2D coordinate access (z defaults to 0).ms[x, y, z]— Full 3D coordinate access.ms[x, y, z] = spectrum— Direct assignment with automatic indexing.
- Special methods
__len__()— Returns the total number of spectra in the collection.__iter__()— Returns an iterator over all spectra in insertion order.__getitem__(key)— Supports flexible indexing by integer or coordinate tuple.__setitem__(key, spectrum)— Assigns spectrum to coordinates with automatic indexing updates.
- Invariants and notes
- Maintains two internal data structures for efficient sequential and coordinate-based access.
- Supports both 2D (x, y) and 3D (x, y, z) coordinate systems.
- Coordinates are automatically extracted and indexed when adding spectra.
- Assignment operations automatically update spectrum coordinates to match the key.
- Example
>>> # Create MS collection
>>> ms = MS()
>>> # Add spectra (user should not load by yourself, use data managers first)
>>> ms.add_spectrum(spectrum1)
>>> # Access by index
>>> spec = ms[0]
>>> # Access by coordinates
>>> spec = ms[10, 20, 0]
>>> # Direct assignment
>>> ms[5, 5, 0] = new_spectrum
>>> # Iteration
>>> for spectrum in ms:
... print(spectrum.coordinates)
output:
[0, 0, 0]
[1, 0, 0]
[2, 0, 0]- Example2
Since the coordinates may be non-contiguous, it is recommended to check the available coordinate range using a mask.
>>> ms_md.load_full_data_from_file()
>>> ms_md.inspect_data()
>>> ms.plot_ms_mask()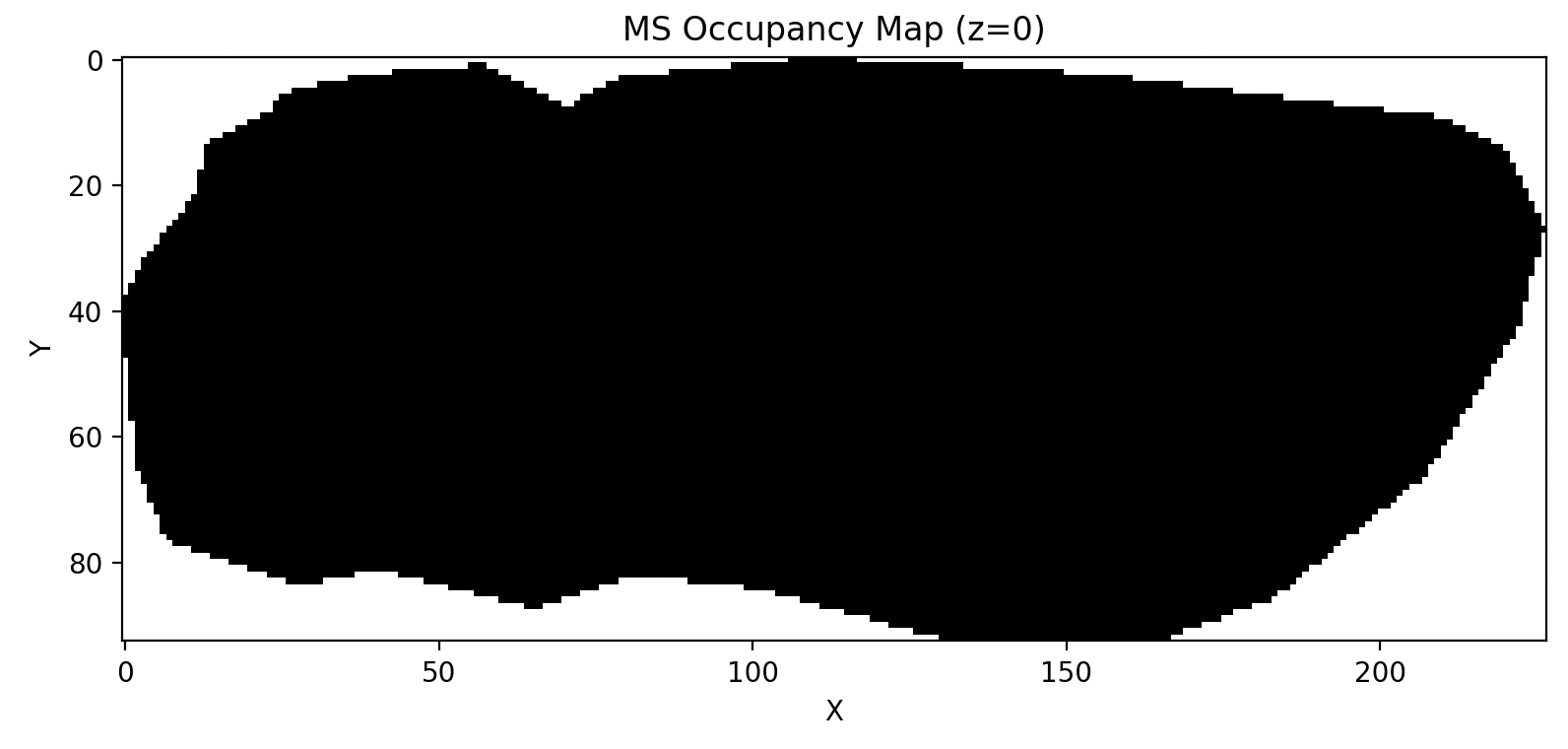
Data Managers
MSDataManager Class (Abstract)
class module.ms_data_manager.MSDataManager(ms, target_mz_range, target_locs, filepath)Abstract base class that defines common configuration and the contract for loading mass spectrometry data into an MS model. This class provides the foundation for concrete data manager implementations that handle different file formats.
- Parameters
ms(MS) — The targetMSdomain model instance to populate with spectra.target_mz_range(Optional[Tuple[float, float]]) — Inclusive range(min_mz, max_mz)to filter peaks. Can beNoneto load all m/z values.target_locs(Optional[List[Tuple[int, int] | Tuple[int, int, int]]]) — A bounding region defined by two coordinates[(x1, y1), (x2, y2)]or 3D equivalents; used to limit loaded spectra. Can beNoneto load all locations.filepath(Optional[str]) — Path to the input data file.
- Methods
get_ms()— Returns the internalMSinstance.load_full_data_from_file()— Abstract method; must be implemented by concrete managers to load data.inspect_data(inpect_num=10)— Logs dataset information including count, sample spectra lengths and ranges.
- Validation
- Ensures
target_locscontains at least two coordinate points when provided. - Validates that
x1 < x2andy1 < y2for bounding box coordinates. - Raises appropriate errors for invalid configuration.
- Ensures
- Invariants and notes
- This is an abstract class and cannot be instantiated directly.
- Concrete subclasses must implement
load_full_data_from_file()method. - Provides common infrastructure for spatial and m/z range filtering.
- Maintains a counter for tracking loading progress.
- Example
# Cannot instantiate abstract class directly
# Use concrete implementations like MSDataManagerImzML
if __name__ == "__main__":
from module.ms_data_manager_imzml import MSDataManagerImzML
from module.ms_module import MS
FILE_PATH = "data/example.imzML"
ms = MS()
# Create MS collection and manager
# Auto-created from parser
with MSDataManagerImzML(ms=ms,target_locs=[(1, 1), (50, 50)],filepath=FILE_PATH) as manager:
# Load data with lazy-loading placeholders
manager.load_full_data_from_file()
print(manager.current_spectrum_num)
output:
10000MSDataManagerImzML Class
class module.ms_data_manager_imzml.MSDataManagerImzML(ms, ms_meta, target_mz_range, target_locs, filepath)Concrete data manager for .imzML files. This class extends MSDataManager to handle ImzML format mass spectrometry imaging data, managing metadata initialization and lazy population of the MS collection with SpectrumImzML placeholders.
- Parameters
ms(MS) — The targetMSdomain model instance to populate with spectra.ms_meta(Optional[MetaDataImzMl | MetaDataBase]) — Metadata wrapper object. IfNone, will be created automatically from the parser.target_mz_range(Optional[Tuple[float, float]]) — Inclusive range(min_mz, max_mz)to filter peaks. Can beNone.target_locs(Optional[List[Tuple[int, int] | Tuple[int, int, int]]]) — Bounding region for spatial filtering. Can beNone.filepath(str) — Path to the.imzMLfile to load.
- Attributes
parser(ImzMLParser) — ImzML parser instance created fromfilepathfor reading spectrum data.meta(MetaDataImzMl | MetaDataBase) — Metadata wrapper bound to the parser, caching commonly used fields like image dimensions and instrument information.
- Inherited Attributes
_ms(MS) — The target domain model to populate.target_mz_range,target_locs,filepath,current_spectrum_num— FromMSDataManager.
- Methods
load_full_data_from_file()— Implements the abstract method to load ImzML data with lazy-loading spectra.get_ms()— Returns the populatedMSinstance.inspect_data(inpect_num=10)— Inherited method for dataset inspection.
- Initialization logic
- Validates and creates an
ImzMLParserfromfilepath. - If
ms_metais not provided, createsMetaDataImzMl(parser=parser)automatically. - If
ms_metais provided but without a parser, injects the parser into the metadata object. - Otherwise uses the provided metadata as-is.
- Validates and creates an
- Invariants and notes
- Only supports
.imzMLfile format; raises error for other extensions. - Uses lazy loading to minimize memory usage—actual spectrum data is not loaded until accessed.
- Metadata is tightly coupled and provides essential information for downstream processing.
- Only supports
- Example
from module.ms_module import MS
from module.ms_data_manager_imzml import MSDataManagerImzML
# Run examples when executing this file directly
if __name__ == "__main__":
FILE_PATH = "data/example.imzML"
ms = MS()
# Create MS collection and manager
# Auto-created from parser
with MSDataManagerImzML(ms=ms,target_locs=[(1, 1), (50, 50)],filepath=FILE_PATH) as manager:
# Load data with lazy-loading placeholders
manager.load_full_data_from_file()
# Inspect loaded data
manager.inspect_data(inpect_num=5)
# Access spectrum (triggers lazy load)
spectrum = ms[40, 13]
print(spectrum.mz_list[:5])
inspect_data output:
INFO: 25-11-10 19:34 202 ms_data_manager - creating ms mask.
INFO: 25-11-10 19:34 102 ms_data_manager - MS meta data:
target_mz_range: None
target_locs: [(1, 1), (50, 50)]
filepath: data/example.imzML
current_spectrum_num: 1910
meta_name: ImzML
meta_version: 1.0
meta_storage_mode: split
meta_centroid_spectrum: None
meta_profile_spectrum: True
meta_max_count_of_pixels_x: 227
meta_max_count_of_pixels_y: 93
meta_pixel_size_x: 100.0
meta_pixel_size_y: 100.0
meta_absolute_position_offset_x: 0.0
meta_absolute_position_offset_y: 0.0
meta_min_pixel_x: 0
meta_min_pixel_y: 2
meta_mask: (93, 227)
INFO: 25-11-10 19:34 115 ms_data_manager - MS information:
MS len: 74749
MS range: 400.0 - 1000.0
MS coord: (0, 38, 0)
max and min mz_list: 1000.0 - 400.0
max intensity: 16.307722091674805
MS len: 74749
MS range: 400.0 - 1000.0
MS coord: (0, 39, 0)
max and min mz_list: 1000.0 - 400.0
max intensity: 17.022132873535156
MS len: 74749
MS range: 400.0 - 1000.0
MS coord: (0, 40, 0)
max and min mz_list: 1000.0 - 400.0
max intensity: 16.470420837402344
MS len: 74749
MS range: 400.0 - 1000.0
MS coord: (0, 41, 0)
max and min mz_list: 1000.0 - 400.0
max intensity: 19.481334686279297
MS len: 74749
MS range: 400.0 - 1000.0
MS coord: (0, 42, 0)
max and min mz_list: 1000.0 - 400.0
max intensity: 18.155176162719727
[400. 400.00802697 400.01605394 400.02408091 400.03210788]Metadata Dependencies
While not part of the three focus modules, MetaDataImzMl (in module/meta_data.py) is tightly coupled to MSDataManagerImzML:
- It stores and exposes commonly used fields (e.g., image dimensions, pixel sizes, instrument model) and computes
spectrum_count_numfromparser.coordinates. - It can be initialized with either a parser or a file path and will extract metadata via
pyimzml.
Data Flow Summary
- Create an empty
MScollection. - Initialize
MSDataManagerImzMLwith theMS, optionaltarget_locs/target_mz_range, andfilepath. - Call
load_full_data_from_file()to populateMSwithSpectrumImzMLplaceholders for spectra within the target region. - When accessing
mz_listorintensityon aSpectrumImzMLspectrum, lazy loading fetches the data from the ImzML file via the parser. - Metadata remains available through
metafor consumers that need image dimensions, pixel sizes, instrument model, etc.
Extensibility Notes
- New managers can subclass
MSDataManagerto support additional formats; adhere to theload_full_data_from_file()contract and, if practical, use lazy-loading spectra likeSpectrumImzML. - Consider using
target_mz_rangeduring load to prefilter peaks for large datasets, or leave it for downstream processing depending on performance needs.
Glossary
- ImzML: An XML-based standard for MSI data storage.
- Spectrum: A pair of arrays
(mz_list, intensity)associated with specific spatial coordinates. - Lazy loading: Deferring actual data reading until first access to reduce memory and I/O overhead.